Input data and parameters
Input
| Analysis date: | Sun Mar 08 00:14:34 GMT 2026 |
| BAM file: | GM12878_REP1.namesorted.bam |
| Counting algorithm: | uniquely-mapped-reads |
| GTF file: | genes.gtf |
| Number of bases for 5'-3' bias computation: | 100 |
| Number of transcripts for 5'-3' bias computation: | 1,000 |
| Paired-end sequencing: | yes |
| Protocol: | strand-specific-reverse |
| Sorting performed: | no |
Summary
Reads alignment
| Number of mapped reads (left/right): | 86,909,417 / 86,888,892 |
| Number of aligned pairs (without duplicates): | 86,778,882 |
| Total number of alignments: | 189,114,475 |
| Number of secondary alignments: | 15,316,166 |
| Number of non-unique alignments: | 26,451,119 |
| Aligned to genes: | 122,500,822 |
| Ambiguous alignments: | 2,422,409 |
| No feature assigned: | 37,740,125 |
| Not aligned: | 12,490,977 |
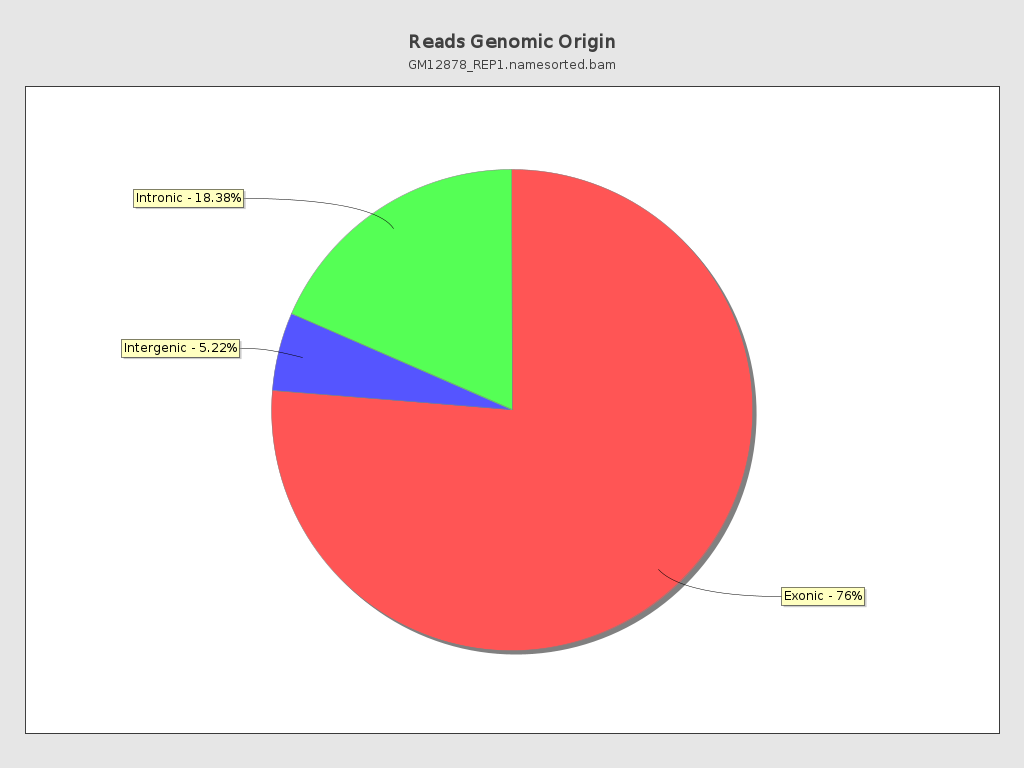
Reads genomic origin
| Exonic: | 122,500,822 / 76.45% |
| Intronic: | 29,428,759 / 18.37% |
| Intergenic: | 8,311,366 / 5.19% |
| Intronic/intergenic overlapping exon: | 6,734,205 / 4.2% |
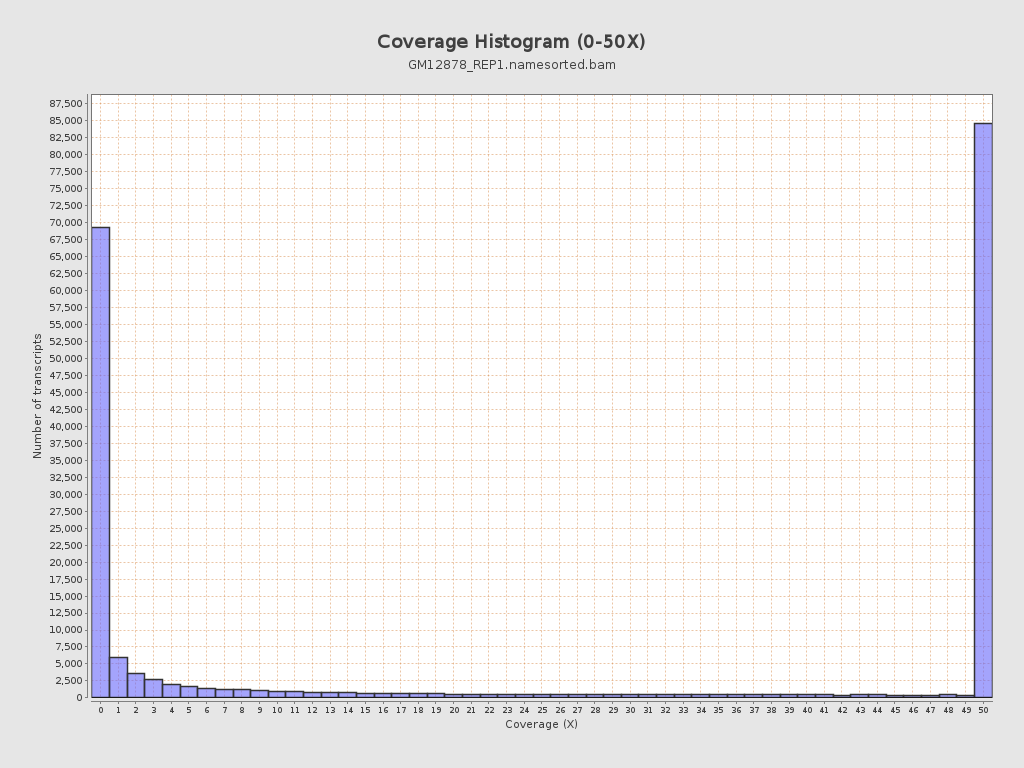
Transcript coverage profile
| 5' bias: | 0.87 |
| 3' bias: | 0.77 |
| 5'-3' bias: | 1.11 |
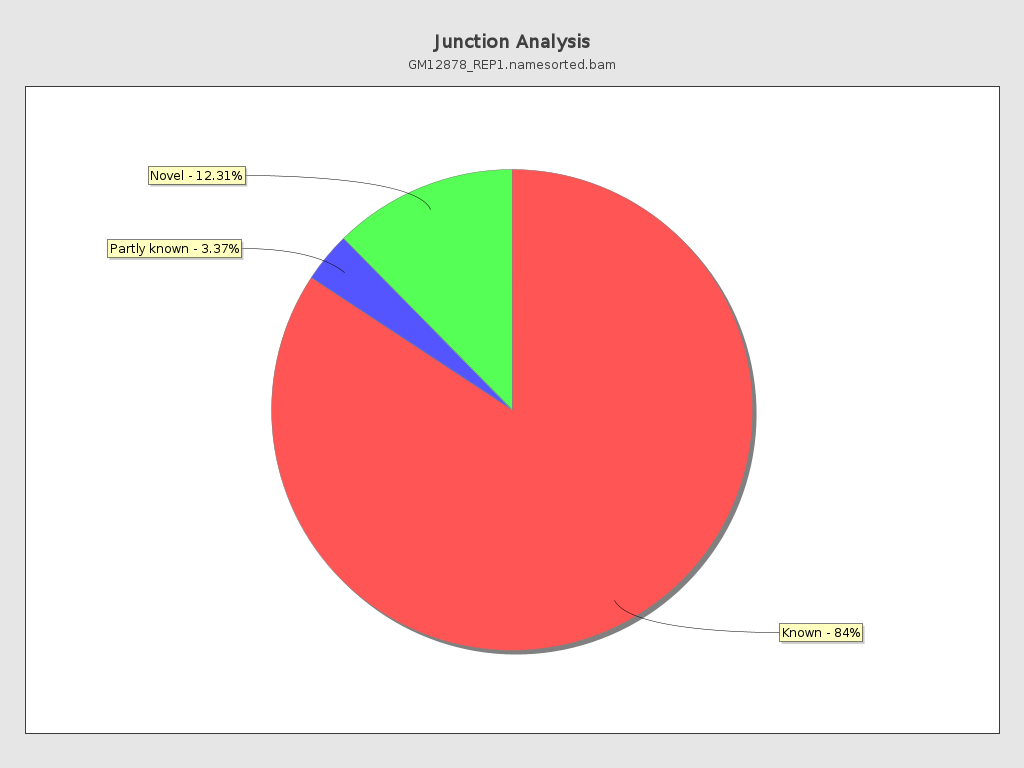
Junction analysis
| Reads at junctions: | 56,119,719 |
| AGGT | 5.41% |
| ACCT | 5.4% |
| AGGA | 3.57% |
| TCCT | 3.29% |
| AGCT | 2.91% |
| AGGC | 2.87% |
| ATCT | 2.86% |
| GCCT | 2.43% |
| CCCT | 2.34% |
| AGGG | 2.3% |
| AGAT | 2.12% |

.png)
.png)
.png)